My case for reproducible research (and some tools)
Tom Ranner
Slides available at tom-ranner.gitlab.io/reproducable-research
Before I forget…
Here you go😀 A "worm" walks as a response to the change of environmental humidity (the tube provides moisture.).https://t.co/ZyD7Dy1pfo. pic.twitter.com/QS7yQ1HYU2
— Sunjie Ye (@SunjieYe) October 23, 2020
What is the problem?
Do I trust my research software?
- Can I trust my code to give the same results when I come back to it weeks, months, years later?
- Can I run my code on which ever machine I want (i.e. laptop, mju, arc, jade, …)?
- Can someone else reproduce the same results as me?
What other people think?
There is concern this is a reproducibility crisis in computational research….
Here are some examples (from https://mikecroucher.github.io/reproducible_ML/)
The famous excel error

The gene excel problem
Gene name errors are widespread in the scientific literature Make Zieman, Yotam Eren and Assem El-Osta
More seriously
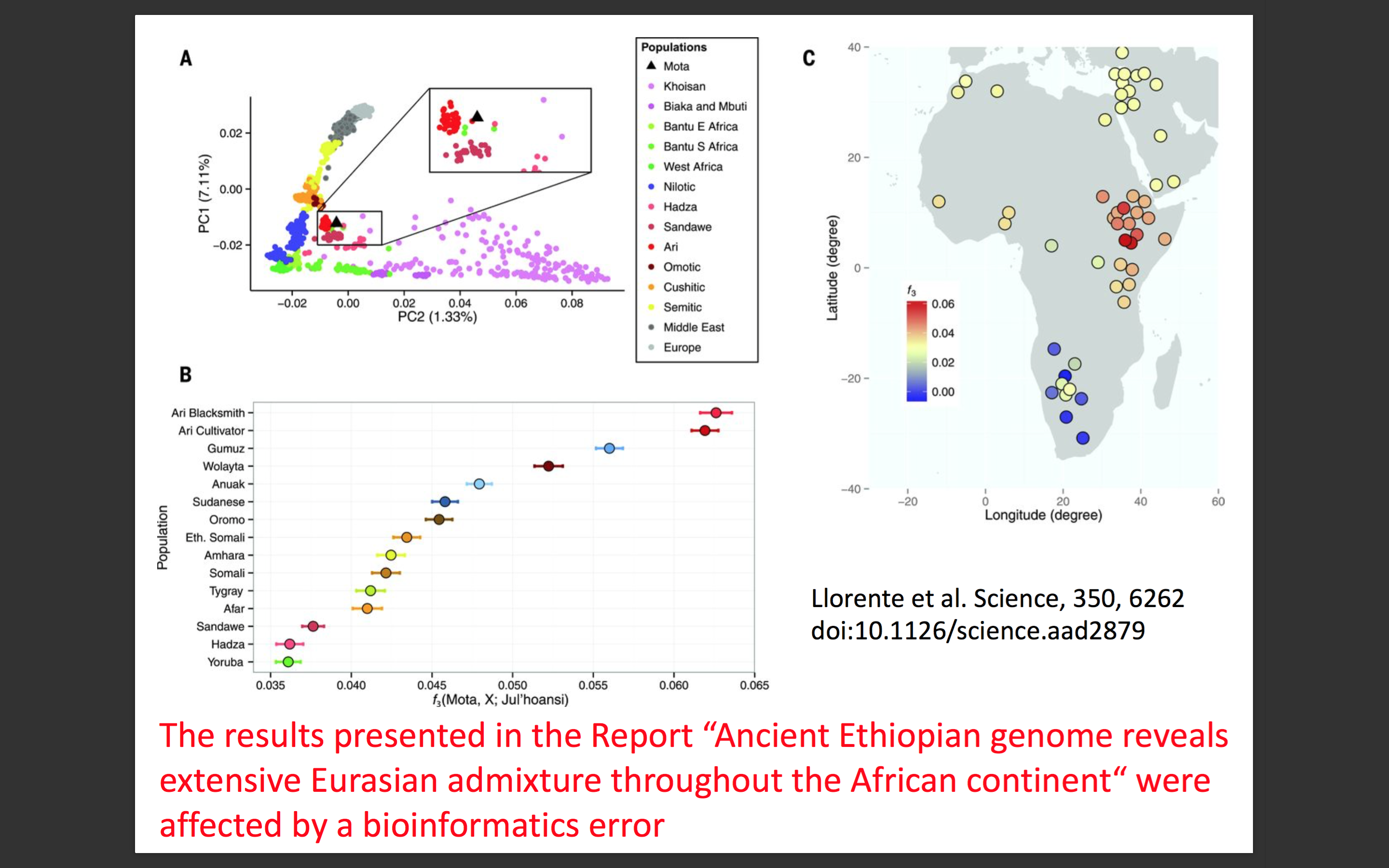
More seriously 2
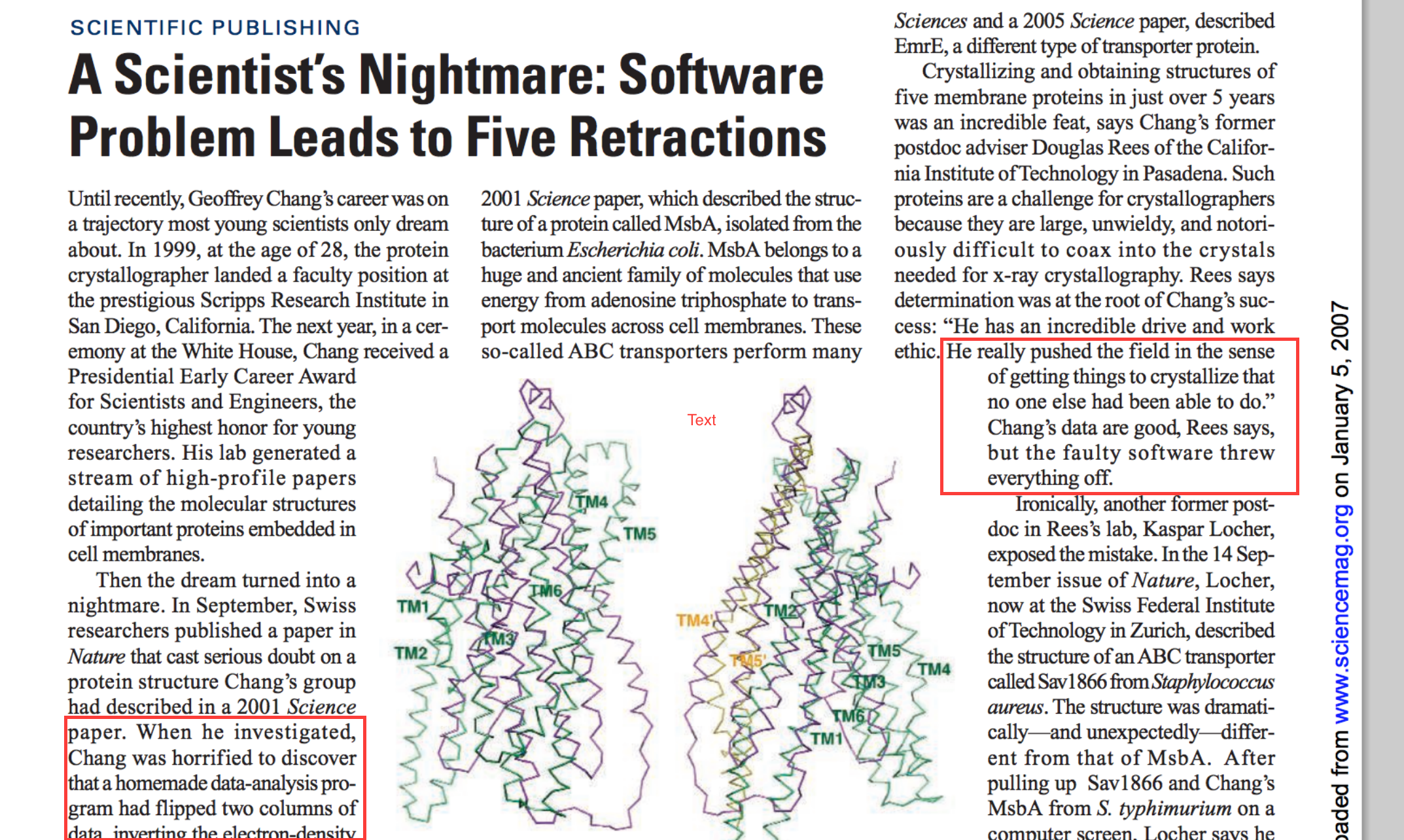

Version problems

Basic good ideas
- clicks aren’t reproducible
- code in high level languages
- source code management (
git) - which version gave you your results? - some way to track environment/dependencies and versions (today)
- ideal case: generate all results, figures, paper, … just by running one command (also today)
- automation is not about saving time
Solution 1: conda (environments)
What is conda?
Conda aims
- Install a specific set of dependencies with well defined versions
- Record dependencies and version for all dependencies
- Isolate environments rather than installing globally
- Different versions of dependencies per project
Conda: how it works?

The details
- Not just for python
- Open source BSD licence (not GPL code)
minicondais a lightweight alternative toanaconda- Can be installed on your computer (windows, mac, linux, hpc, …) without admin rights
Let’s have a go!
- creating and activating a new conda environment
- installing packages
- saving environment to file
- removing an environment
- installing from file
Solution 2: singularity
What is singularity?
Wait - what’s a container?

A container is a standard unit of software that packages up code and all its dependencies so the application runs quickly and reliably from one computing environment to another.
Singularity aims
- Available for most operating systems
- A mechanism to send the computer to the data
- Solve the problem of getting code running on another computer by sending the computer
- Singularity is aimed at the scientific community to run mainly using HPC
- Can use docker images - for example stored at DockerHub
- Also singularity recipes and images
Some language
Image is a blueprint. It is immutable.
Container is an instance of an image.
Dockerfile/Singularity recipe is a recipe which creates a container based on an image and potentially applies small changes to it
Pros
- Allows for seamless moving workflows across platforms
- Lightweight solution (c.f. virtual machines)
- Eliminates the works on my machine problem
- Very straightforward dependency management
- Doesn’t require root access to run (requires root to build)
Cons
- There are potential security issues
- where did you get your image from?
- Can be used to hide away software install problems and thereby discourage good software development practices
- Why use
cmakewhen you know the path of all dependencies?
- Why use
Let’s have a go
pulling containers (potentially slow)- getting a shell in a container
- what files are here already?
- some of the magic
- what a recipe looks like
Solution 3: snakemake
What is snakemake?

https://snakemake.github.io/ | https://snakemake.readthedocs.io/en/stable/
Snakemake’s aims
- A special tool to reproduce a number of steps in a computational workflow
- Replacement for clicking lots of buttons in a GUI, bash script, makefile, …
- Gentle learning curve
- Cross platform available via conda (bioconda channel)
- Heavily used in bioinformatics but completely general
How it works

Pros and cons
- Designed for scientific workflows including HPC or kubernetes
- Integrates really well with conda and singularity
- Can only really be set up once you know what the pipeline is
- Requires all tasks to be on one machine
- (I’ve not used this tool seriously so far but it seems useful…)
Let’s have a go
- What’s a
snakefile? - How to generate the dag image?
Other solutions or helpers
Other ideas
git annexorgit lfsfor data storage- testing - gives you confidence to make changes to code
- use well established libraries
- code review
- continuous integration/continuous deployment (gitlab, bitbucket, sourcehut)
